Visualization & Analysis¶
Inscoper I.S. provides tools for visualizing data during and after acquisition.
Visualization During Acquisition¶
The Visualization tab allows for real-time monitoring of the acquisition sequence.
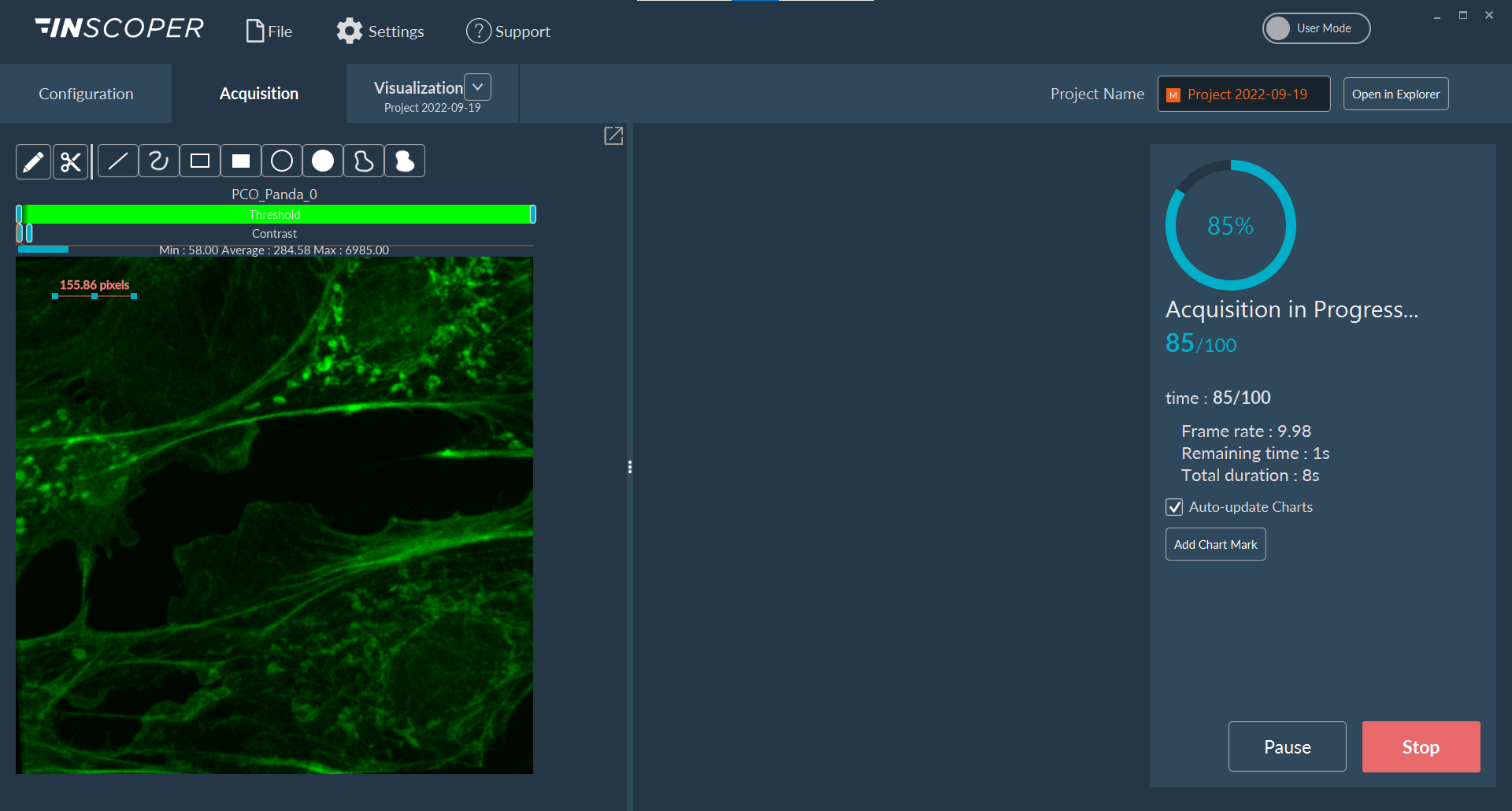
The interface is divided into three functional areas:
- Left Panel: Displays the live, incoming images from the current acquisition sequence.
- Center Panel: Renders corresponding graphical data (e.g., intensity evolution or statistical plots), depending on the configured Data Processor pipeline.
- Right Panel: Provides acquisition progress metrics and system controls to pause or abort the active sequence.
Large Area Tiling
During extensive tiling acquisitions, the left panel dynamically updates to display the full macroscopic mosaic as individual tiles are acquired and aggregated into the global view.
Visualization After Acquisition¶
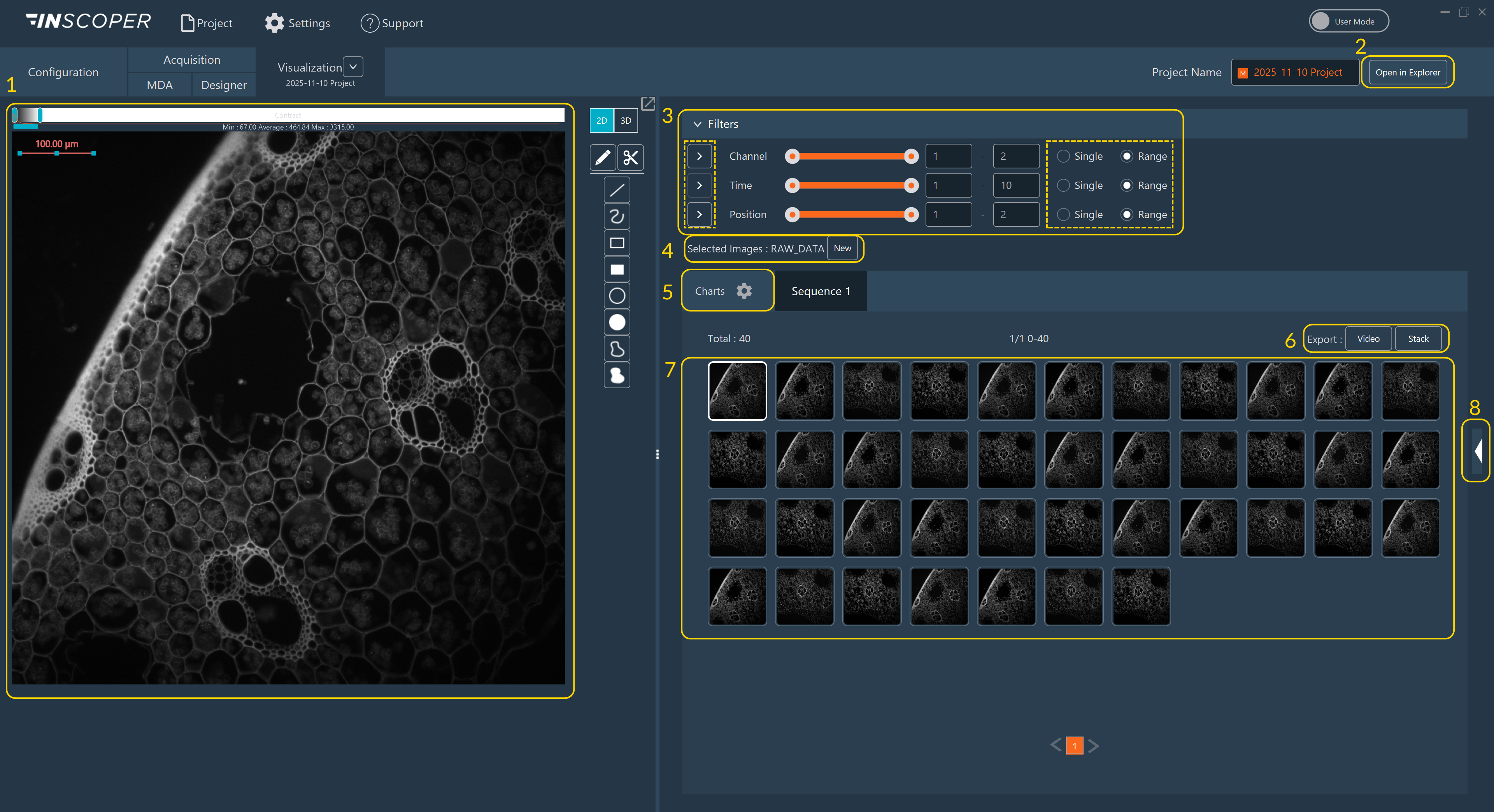
- Image Canvas: View images in the central viewer.
- Open Directory: Navigate directly to the local dataset storage path (applicable only if data is saved to disk).
- Filters: Use dimension-based filters to search your dataset. Select individual images or full groups. Click the Play button adjacent to a dimension name to animate the sequence.
- Data Processing: Select output images for post-acquisition processing.
- Graphics Mode: Switch viewing modes to visualize quantitative data charts.
- Export Options: Save the currently viewed sequence as an MP4 Video or a TIFF Stack.
- Thumbnail Gallery: Browse miniature representations of all acquired images.
- Metadata Access: Reveal the hardware and experimental parameters tagged to the images.
3D Viewer¶
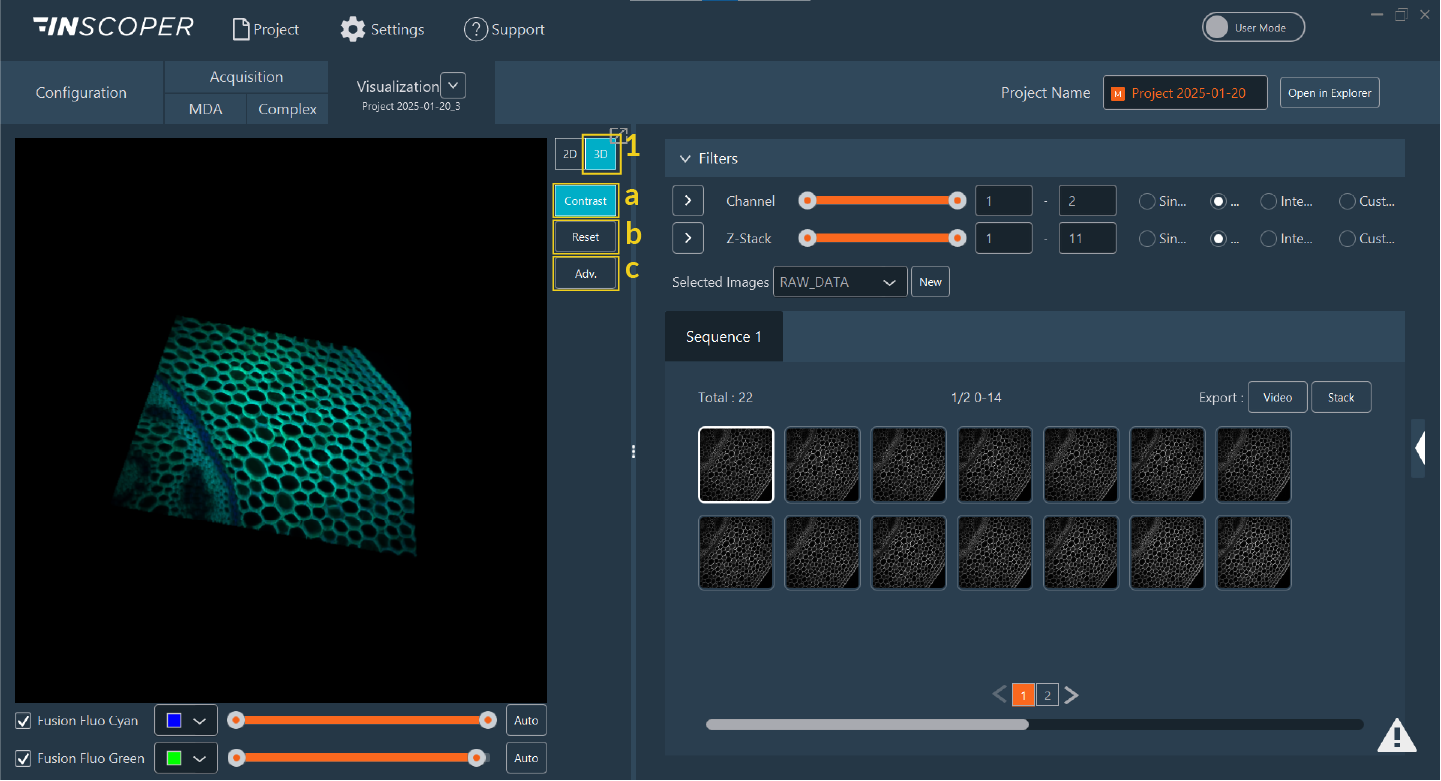
- Click the 3D button to launch the volumetric viewer.
- 3D visualization canvas.
-
Adjust rendering options and parameters:
- Contrast: Display the LUT and adjust the contrast bar below the 3D view.
- Reset: Restore the default 3D camera view.
-
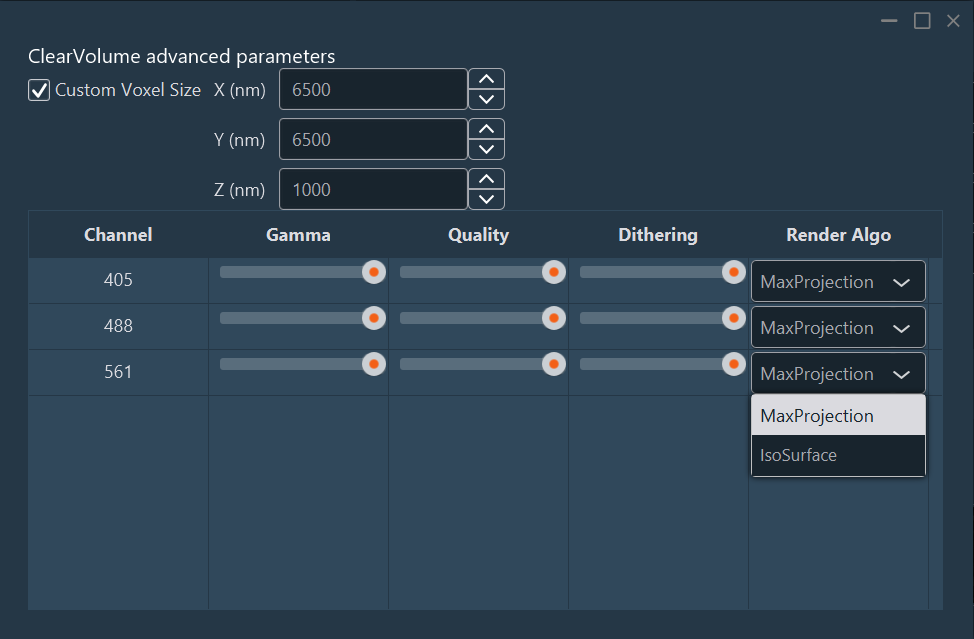
Advanced: Access detailed rendering parameters per channel:
- Voxel Size (nm).
- Gamma: Apply a non-linear gamma curve to adjust contrast.
- Quality: Adjust rendering quality for navigation.
- Dithering: Reduce banding/aliasing artifacts.
-
Render Algorithms:
- MaxProjection: Displays only the voxels with the maximum intensity encountered along the viewing ray.
- IsoSurface: Renders a 3D surface connecting points of equal intensity value.
Interacting with Graphics¶
Interact with charts by adjusting their appearance, adding time markers, or exporting raw data. Hover over a graph to access its toolbars.
- Pan: Click and drag the mouse wheel (middle click) to pan across the graph.
- Zoom: Scroll the mouse wheel to zoom in and out. Click and drag to select a region to magnify.
- Jump to Image: Left-click a data point on the graph to view the corresponding image.
- Reset View: Right-click within the graph to restore the default axis scaling.
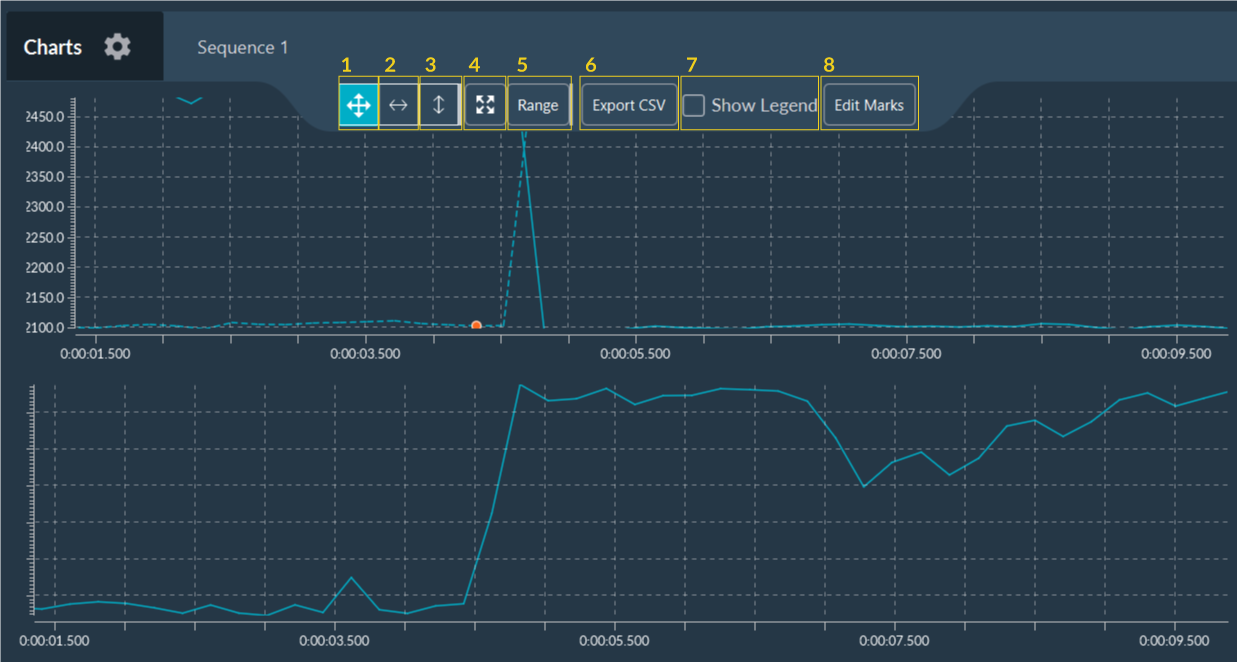
Graph Tools¶
- Enable the XY zoom mode
- Enable the X zoom mode
- Enable the Y zoom mode
- Reset zoom or enable auto-ranging.
- Manually override the axis ranges.
- Export the raw graph data to a
.csvfile. - Toggle curve legends.
- Add Markers: Annotate the timeline with event markers. Markers can be exported to a
.csvfile.
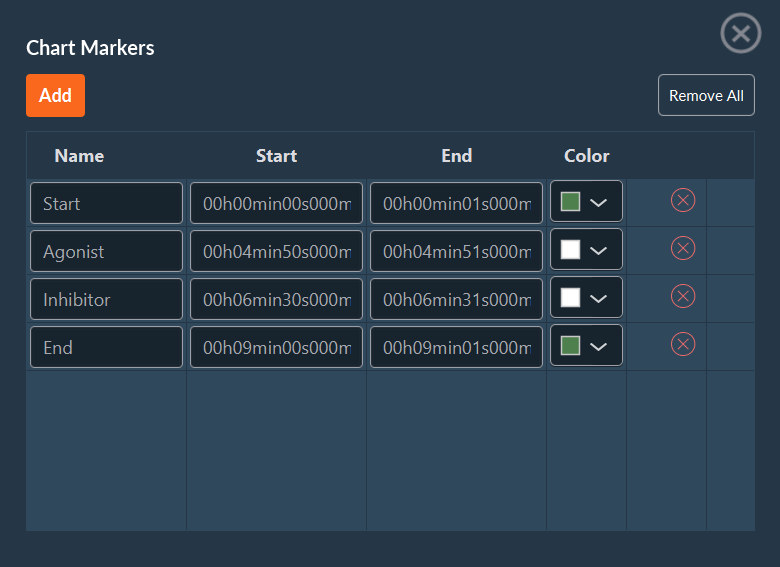
In this window you can add information about your experiment as markers. These events, which are fully customizable, can be associated with the acquisition itself (start, pause, end), external events (addition of an inhibitor, medium supplementation), or others. These markers can be saved in a .csv file and reused at any time.
Image Processing¶
Apply post-acquisition data processing from the Visualization tab.
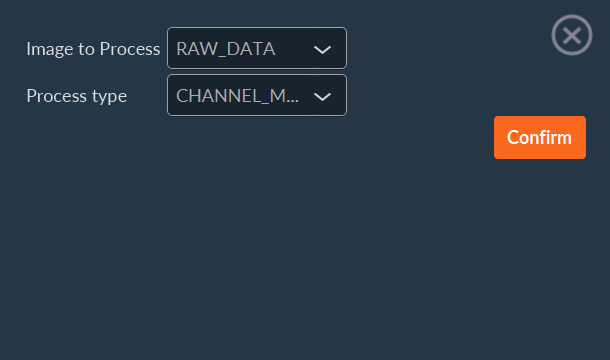
- Select the target sequence from the image drop-down list.
- Choose the desired data processor.
- Click Confirm to execute.
Example: Tiling Processing¶
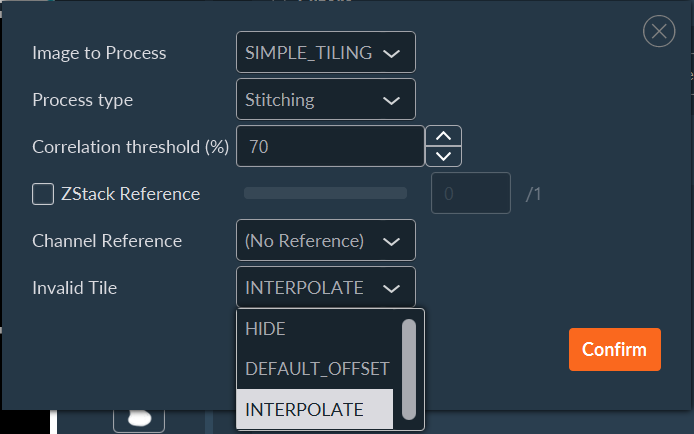
- Select the source sequence.
- Select the Tiling process type.
- Set the cross-correlation threshold (percentage) for image stitching.
- (Optional) Apply a Z-Stack reference, specifying which focal plane should drive the 2D stitching calculation.
- (Optional) Select a specific reference channel to calculate the stitching coordinates.
-
Define the fallback behavior for uncorrelated (invalid) tiles:
- HIDE: Discard the tile from the final image.
- DEFAULT_OFFSET: Place the tile based solely on its recorded mechanical stage coordinates.
- INTERPOLATE: Interpolate the tile's position based on the calculated offsets of surrounding valid tiles.
-
Click Confirm.
Data Export¶
Export sequences as MP4 videos or multi-dimensional Tiff stacks via the Export menu (Toolbar Item 6).
Video Export¶
Use dimension filters to isolate the image sequence, then click Video from the Export dropdown.
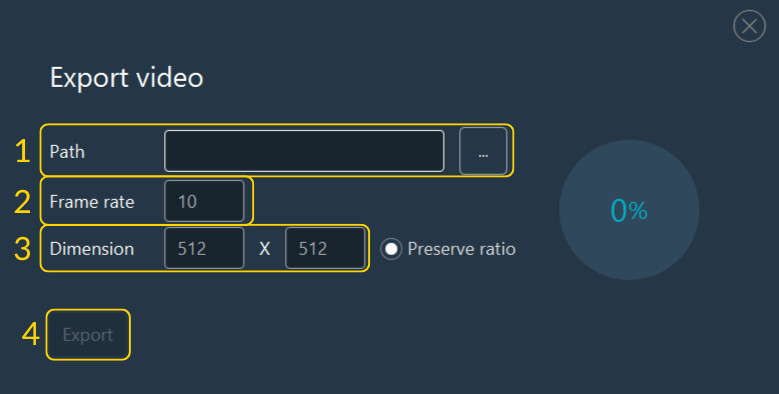
To export a video:
- Select the destination file path.
- Verify the sequence to export.
- Select the video format/codec.
- Click Export.
Stack Export¶
Use dimension filters to isolate images, then click Stack from the Export dropdown to generate a multidimensional TIFF.
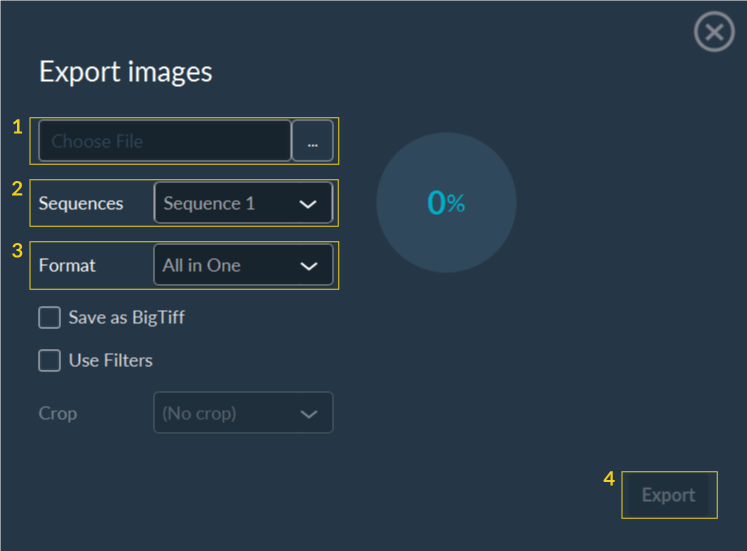
To export a stack:
- Select the destination file path.
- Verify the sequence to export.
- Select the data format.
- (Optional) Check Save as BigTiff if the expected file size exceeds 4GB.
- (Optional) Check Use Filters to apply your visualization filters to the export.
- Click Export.
Metadata Access¶
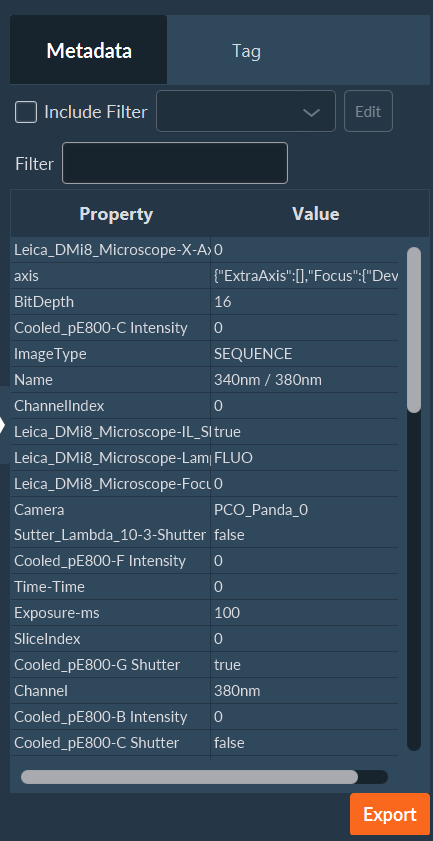
On the right side of the window there is a white triangle. You can click on it to access all the metadata. In this tab, you can access all the metadata, including the camera, light source or microscope settings; a search bar and some filters are available to facilitate the search for some specific parameters. This list can also be exported, if necessary, by clicking on the Export button located in the lower right part of the screen. All metadata are bio-format compatible.
Semi-automated feedback microscopy feature¶
The software supports feedback microscopy workflows. For example, you can acquire a low-magnification overview scan, tag regions of interest, and export those coordinates into a new, high-magnification acquisition sequence.
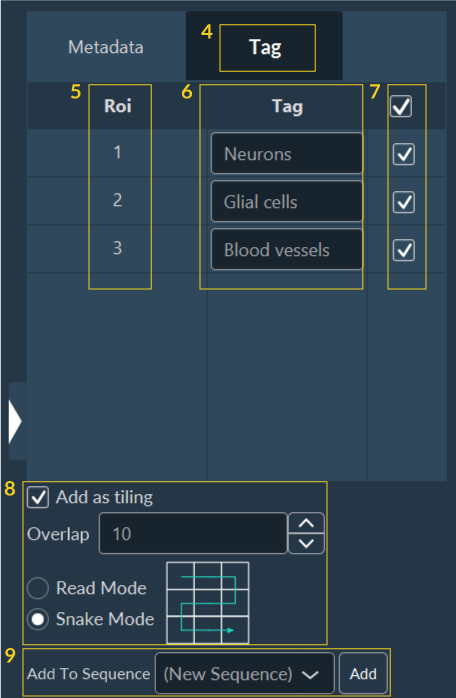
To use this feature:
- Acquire an overview image (e.g., a Tiling scan).
- Use the ROI tools to draw regions around structures of interest.
- In the Visualization tab, click the white triangle on the right edge to expand the side panel.
- Navigate to the Tag sub-tab.
- This lists all drawn ROIs. You can manage them directly from this list.
- (Optional) Assign descriptive names/tags to each ROI.
- Select the ROIs you wish to target.
- If the new targets require their own sub-tilings, configure those spatial parameters here.
- Choose to inject these new coordinates into a fresh sequence or append them to an existing one.
Smart Microscopy
Use this workflow for:
- A low-magnification pre-scan followed by targeted high-resolution imaging.
- A brightfield pre-scan, applying fluorescence imaging to identified structures to reduce global phototoxicity.